Abstract
Introduction:
The endometrial cancer (EC) is the seventh most common malignancies worldwide among females with good prognosis in early stages of the disease. The CpG Island in the promoter region of tumor-suppressor genes are frequently methylated in various types of human cancers. In the present study, we investigated the methylation pattern in promoter region of RASSF1A and RASSF2A genes in Endometrial cancer patients in Iranian women to identify correlations among promoter hypermethylation, disease risk and clinicopathological parameters.Methods:
28 patients and 22 healthy controls were studied. Isolation of genomic DNA from FFPE and peripheral blood was performed and Methylation-Specific PCR (MSP) was applied for analysis of the promoter CpG methylation status of RASSF1A and RASSF2A genes in the studied population.Results:
A significant difference was found among the study groups and the presence of promoter CpG hypermethylation status in the RASSF1A (P = 0.0321) and RASSF2A (P = 0.0003) genes. RASSF1A, and RASSF2A gene promoter methylations were present in 53.57% and 42.85% of EC samples when compared to those in the controls with 31.81% and 9.09% respectively. Furthermore, methylation status between tissue and blood samples of RASSF1A, and RASSF2A genes was not significant (P = 0.49 and 0.09 respectively). Our results indicated a corollation between ages, menosososal state and tumor grade with RASSF1A, and RASSF2A promoter methylation.Conclusions:
In our study, Hypermethylation of both RASSF1A and RASSF2A genes are important events in carcinogenesis of endometrial cancer. Epigenetic alternations may have diagnostic value for early diagnosis and better clicinal management of susceptibility to endometrial malignancies.Keywords
1. Intruduction
Endometrial cancer (EC) is the major cancer of the female genital tract in the United States and the fourth most common cancer among women after breast, lung, and colorectal cancer (1), and the risk of its develpment in women is about 2.6 percent (2). Aproximately 90% of cases are sporadic and 10 % are hereditary type. Development of endometrial cancer is a multistage process of activiation of proto-oncogenes and inactivation of tumor suppressor genes, according to histopathology, cell biology and clinical course is classified into two main types: type I carcinomas, slow endometrioid diffrentiation and associated with unopposed estrogen exposure; type II carcinomas mainly composed by serous and clear cells and follow estrogen unrelated pathways (3). Most endometrial cancers are the type 1 that often result from an endometrial background hyperplasi; in contrast, type 2 is less common and often is detected by non endometrial tissues and during the clinical invasion. The first and most common symptom of endometrial cancer is abnormal bleeding from the vagina that should be taken seriously. Risk factors for EC include: obesity, anovularity states, infertility, early onset of menstruation, late menopause, nulliparity, and exposure to estrogen therapy that would promote development of endometrial hyperplasia with or without atypia which is a possible precursor lesions of EC (4-6).
Genetic and epigenetic factors play important roles in the development of endomentrial cancer. In previous reports, genetic mutations of oncogens such as p53, K-RAS and PTEN have been studied. However, recently it is known that epigenetic changes such as methylation, histon deacetylation, or micro RNA expressiorn can also affect the molecular biology of endometrial lesions on lining of the uterus (7, 8). Epigenetics refer to heritable changes in gene activity and expression without changing the DNA sequence by pharmocological agents is dynamic and reversible. The most common epigenetic event is aberrant DNA methylation in CpG dinucleotides by three types of DNA methyl transfrase including DNMT3A, DNMT3B and DNMT1, in which methyl group (CH3) covalently attatches to cytosine, and creates 5 methyl cytosine (9). Promoter hypermethylation is associated with silencing of tumor suppressor genes involved in cell cycle regulation, DNA repair, cell-cell interaction, steroid receptors, apoptosis and angiogenesis (10). Thus, in the present study, promoter hypermethylation of two key genes involved in regulatin of MAP kinase signaling pathway was investigated.
RASSFIA, the first member of the RASSF family, as a tumor suppressor gene, located on chromosome 3p21.3, contributes to numerous cellular functions. This protein regulates apoptosis in two ways: in the first case, attached to MST1 and Nore1 proteins, resulting complex, detecting apoptosis signals, and thereby controlling the RAS signaling pathway. In the second state, RASS1FA regulates apoptosis by MOAP1 interaction (apoptosis modulators and related bc12 family) which is followed by BAX and caspase dependent expression (11-14). RASSF1A is thought to be involved in cell cycle regulation through a direct connection with cdc20 (a negative regulator of APC), proliferation, cell mobility and adhesion (15). Also involved in regulation of the cyctoskeleton dynamic, microtubules spindle apparatus and centrosome during metaphase and microtobule stability (16). This protein inhibits passing G1 phase to S, through corporation with P120E4F (a known protein related to RB, P53 and p14ARF) (17, 18). As well as inhibition of cycline D1 complex accumulation through the JNK kinase pathway and suppression the activity of AP1, cell cycle arrest in G1/S phase. Also RASSF1A induces cell cycle G1 arrest by increasing the activity of a known transcription factor, P120E4F, which is involved in cycline A2 transcription,and simulates the ubiquitination of MDM2 protein (negative regulator of P53), so it increases the P53 protein activities (19-21).
RASSF2A, the second member of RASSF family, is localized on human chromosome 20p13 (22). Studies have described RASS2FA nuclear localization sequence (NLS) at the N terminal part, is required for full the tumor suppressor activity of the protein (23). Suppressive properties, due to its ability for apoptosis regulation and cell cycle progression, it has been shown that RASS2A is effective in stability and mobility of microtubules (24), also it is known as a potential suppressor of apoptosis, cell cycle arresting and increased cell-death by k-RAS inhibiting activiteis. It has been shown that RASSF2A connects with MST1 with a manner beyond simple communication of protein-protein interaction, and plays an important role in MST1 regulation (25, 26). RASSF2A has a distinct role in the regulation of the MST2 performance. Interestingly, RASSF2A overexpression leads to increased levels of MST1, and provides MST1 protection. RASSF2A appears to be a substrate for MST1 and MST2 and co-expression of either kinase with RASSF2 relocalises RASSF2 from the nucleus to the cytoplasm in a manner dependent on kinase activity (23). RASSF2A has also been shown to be frequently inactivated by promoter methylation in a wide range of tumor types.
While epigenetics refers to broad changes in several types of malignancies, including gynecological. We focused on the role of DNA methylation in relation to endometrial carcinogenesis. We aimed to investigate the aberrant methylation of CpG islands within the promoter regions of two tumor-suppressor genes RASSF1A and RASSF2A, in the tumor tissue and blood samples from patients with endomentrial cancer in comparison with normal tissue, in order to define the frequency of the epigenetic alterations and to determine the possible impact on the disease histological pattern.
2. Methods
2.1. Characterization of Clinical Specimens
A Number of 28 tumor and 22 normal tissues from women with endometrial carcinoma were collected at the time of Hysterectomy by a physician obstetrics and gyneocogy, and were identified through ultrasound. Meanwhile, 26 blood samples from related patients have been collected. Written informed consent was obtained from all study participants (ZUMS.REC.1395.141). Formalin Fixed Paraffin Embedded tissues were kept in laboratorory tempature and prior to DNA extraction hematoxylin- eosin staining was performed and histopathology diagnosis of endomentrial carcinoma was approved by pathologist (Figure 1). Blood samples were collected in tubes containing EDTA at - 20°C temperature for long-term storage. Among the 28 endometrial cancers, the average age of patients was 65.56 years (range 32 - 76 years). All the women with abnormal bleeding were refered to a doctor, 18 cases tumor grade G1 and 10 patients grade had G2 and G3. Most tumors in stage IA endometrioid carcinoma were discretional and only 4 patients had papillay serous type. Myometrial invasion in 15 subjects less than 50% and in 10 cases more than 50 percent and the remaining 3 were not myometrial invasion. Also 9 people were cases of metastasis to other locations and 19 did not have metastasis. 11 patients were with diabetes, 8 patients with high blood pressure, 4 patients with low thyroid and 3 patients had hypelipidemind. 19 of these patients were added to Verne, one of them for 2.5 years of hormonr use had.
Microdissection of Endometrial Cancer Cells, (H & E, × 100), Neoplastic Cells with Enlarged and Coading Nuclei, Which Occupied Most of the Cell Volum, Seems to Have Different Size in Comparision with Normal Cells, Abundant Clear Cytoplasm

2.2. DNA Extraction
Bisulfite modification and Methylation- specific PCR: 10 µm- thick sections from each tissue blocks were deparaffinized and genomic DNA was purified by the DNA extraction kit (Qiagen, USA), then 1 - 2 µg of extracted DNA was used for sodium bisulfite treatment (Qiagen, USA). Treatment of genomic DNA with sodium bisulfite converts unmethylatedcytosines (but not methylated cytosines) to uracil, which is then converted to thymidine during the subsequent PCR step, giving sequence differences between methylated and unmethylated DNA. Methylation status was determined by methylation specific PCR (MS-PCR). The primer sequences for RASSF1A and RASSF2A genes are listed in Table 1. The reaction volumne of 20 µL contained 100 ng bisulfite- modified DNA, 10 mL 2X Taq premix (master mix) [Pars Tous, Iran], 0/5 µM of each primer and 7 mL of deionized water was used. Amplification of two genes were performed under the following condition: initial denaturation at 95°C for 5 minutes, followed by 35 cycles, at 95°C for 45 seconds (denaturation), at 56°C for 45 seconds (annealing), at 72°C for 45 seconds (elongation), with a final extension for 5 minutes at 72°C. Genomic DNA not treated for bisultite modification and water blanks without added DNA were included as negative controls in each assay. PCR products were analyzed on 2% agarose gel containing safe staine. The PCR for all sampels demonstrating methylation was repeated at least once.
Sequences of Primers, Annealing Temperature and MSP Product Size
| Gene | Primer Sequences 5’ - 3’ | PCR Product Size, bp | Tm, °C |
|---|---|---|---|
| RASSF1A (FM) | GTGTTAACGCGTTGCGTATC | 93 | 58 |
| RASSF1A (RM) | AACCCCGCGAACTAAAAACGA | ||
| RASSF1A (FU) | TTTGGTTGGAGTGTGTTAATGTG | 105 | 55 |
| RASSF1A(RU) | CAAACCCCACAAACTAAAAACAA | ||
| RASSF2A(FM) | GTTCGTCGTCGTTTTTTAGGCG | 89 | 56 |
| RASSF2A(RM) | A AAAACCAACGACCCCCGCG | ||
| RASSF2A(FU) | AGTTTGTTGTTGTTTTTTAGGTGG | 121 | 57 |
| RASSF2A(RU) | AAAAAACCAACAACCCCCACA |
2.3. Statistical Analysis
Association between methylation frequencies of RASSF1A and RASSF2A genes with endometrial carcinoma (tissue and blood samples) and clinicopathologic parameters was statistically analyzed using chi- square (X2) and Fisher exact test. Logistic regression analysis was performed to estimate odds ratios (OR) and 95% confidence intervals (CI). Values of P < 0.05 were considered to indicate a statistically significant difference. All analyses were performed using the SPSS 20 statistical software.
3. Results
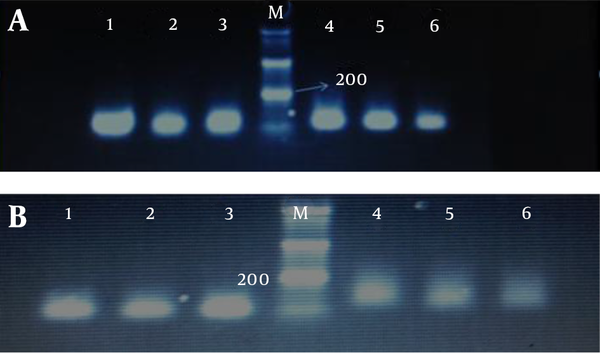
We determined promoter hypermethylation frequency of tumor suppressor genes RASSF1A and RASSF2A in endometrial carcinoma and blood samples of patients and compared the results with normal tissues Also the relationship between hypermethylation of genes with clinical and pathology parameters was evaluated. After treatment of the extracted DNA with sodium bisuflite to convert non-methylated cytosines to uracil, MSP analysis in the study of genes by methylated and non-methylate primers, respectively show 105, 93 bp and 121, 89 bp bands (Figure 2).
MPS Analysis of A, RASSF1A and B, RASSF2A Genes
3.1. RASSF1A and RASSF2A Genes Methylation Distribution
Methylation frequencies of RASSF1A and RASSF2A genes in tumor and normal tissues and blood samples have been shown in Table 2. Promoter methylation of RASS1FA and RASSF2A were more frequent (53.57%, 42.85%) in tumor tissues against those of normal tissues (31.81%, 9.09%). And there was a significant difference (P = 0.03, P = 0.003) respectively. Our results indicate that patients who show hypermethylation of RASS1FA and RASS2FA genes have significantly increased risk of developing endomethrial carcinoma. In contrast there was no statistical association observed beetween methylation status of blood and tissue samples of RASS1F1 gene (P = 0.49); therefore, blood sample can be introduced as a prognostic marker for non-invastive early detection of EC. Our results confirmed that there was no statistically significant difference in promoter methylation analysis of RASSF2A between tissue and blood sample of patients (P = 0.09).
Comparison the Results of Promoter Hypermethylation of RASSF1A and RASSF2A Genes in Tumor and Normal Tissues and Blood Samplesa
| Gene | Sample, No. | Methlyted | Hemi-Methlyted | Non-Methlyted | OR | 95%CI | P Value |
|---|---|---|---|---|---|---|---|
| RASSF1A | Healthy tissue (22) | 7 (31.81) | 9 (40.90) | 6 (27.27) | |||
| RASSF1A | Patients tissue (28) | 15 (53.57) | 11 (39.28) | 2 (7.14) | 2.4957 | 1.0812 - 5.7608 | 0.0321 |
| RASSF1A | Patients blood (26) | 17 (65.28) | 7 (26.92) | 2 (7.69) | 0.7333 | 0.3010 - 1.7864 | 0.4948 |
| RASSF2A | Healthy tissue (22) | 2 (9.09) | 10 (45.45) | 10 (45.45) | |||
| RASSF2A | Patients tissue (28) | 12 (42.85) | 15 (53.57) | 1 (3.57) | 4.9160 | 2.0957 - 11.5316 | 0.0003 |
| RASSF2A | Patients blood (26) | 3 (11.53) | 22 (84.61) | 1 (3.84) | 1.9664 | 0.8938 - 4.3263 | 0.0928 |
3.2. Association with Clinopathological Factors
Based on the obtained results through statical analysis, definitely diabetes and obesity are important risk factors for endometrial cancer incidence (P = 0.016, P = 0.026), but hypothydoidism, hypertension, hypercholesterolemia do not seems as risk factors. When correlated with clinical data, there was marginal association between methylation of RASSF1A gene (tissue and blood sampels) and menopause (P = 0.006). In the case of menopause at the age of 50 years and older, and promoter methylation of RASSF2A gene in tissue, 24 samples have methylation status, so our results have confirmed a significant correlation between incidence of carcinoma at the age of 50 years and older with methylation of RASSF2A gene in the tissue samples (P = 0.013). In regards to the histopathological variables and CpG methylation in the promoter region of both tumor suppressor genes, we found a weak correlation between the RASSF2A gene methylation in tumor tissues and the tumor grade (p=0.09). There was no significant difference in tumor stage, higher tumor grade, type of endometrial carcinoma, myometrial invasion, positive metastasic involvement of pelvic lymph nodes and overweight among methylation of endometrial tumor and normal tissues (P > 0.05) (Table 3 - 5).
| Clinical Parameters | RASSF1A Methylation in Blood | P Value | RASSF1A Methylation in Tissue | P Value | RASSF2A Methylation in Blood | P Value | RASSF2A Methylation in Tissue | P Value | |
|---|---|---|---|---|---|---|---|---|---|
| Age | < 50 | 4 (15.4) | 0.530 | 2 (8.0) | 0.180 | 4 (15.4) | 1 | 3 (11.1) | 0.013 |
| ≥ 50 | 20 (83.3) | 23 (10.7) | 22 (84.5) | 24 (100) | |||||
| Menopause | Before | 4 (15.4) | 0.530 | 2 (8.0) | 0.006 | 4 (15.4) | 1 | 3 (11.1) | 0.013 |
| After | 20 (83.3) | 23 (10.7) | 22 (84.5) | 24 (100) | |||||
| Grade | G1 | 16 (66.7) | 0.618 | 17 (63.0) | 0.750 | 17 (65.4) | 1 | 18 (100) | 0.092 |
| G2 | 4 (16.7) | 5 (18.5) | 4 (15.4) | 4 (80.0) | |||||
| G3 | 4 (16.7) | 5 (18.0) | 5 (19.2) | 5 (100) | |||||
| Stage | IA | 13 (56.5) | 0.714 | 16 (94.1) | 0.644 | 15 (60.0) | 0.221 | 17 (63.0) | 0.183 |
| IB | 2 (8.7) | 2 (100) | 2 (8.0) | 2 (7.4) | |||||
| II | 3 (13.0) | 3 (75.0) | 3 (12.0) | 3 (11.1) | |||||
| AIII | 3 (13.0) | 3 (100) | 3 (12.0) | 3 (11.1) | |||||
| BIII | 2 (8.7) | 2 (100) | 2 (8.0) | 2 (7.4) | |||||
| Type | Endometrioid carcinoma | 21 (87.5) | 0.595 | 24 (96.0) | 0.724 | 32 (88.0) | 0.713 | 24 (96.0) | 0.724 |
| Serous papillary | 3 (12.5) | (100)3 | 3 (12.0) | 3 (11.1) | |||||
| Invasion | Invading not meyometrial | 2 (12.5) | 0.164 | 2 (7.7) | 0.112 | 3 (11.5) | 1 | 3 (11.1) | 0.393 |
| Invasion less than 50% | 12 (50.0) | 15 (57.5) | 14 (53.8) | 15 (55.6) | |||||
| Invasion more than 50% | 10 (41.7) | 9 (34.6) | 9 (34.6) | 9 (33.3) | |||||
| Metastasis | Yes | 9 (39.1) | 0.180 | 9 (32.1) | 0 | 8 (32.0) | 0.497 | 8 (29.9) | 0.139 |
| No | 14 (60.9) | 19 (67.9) | 17 (68.0) | 19 (70.4) |
Comparison of Clinical Features and Methylation of RASSF1A and RASSF2A Genes
| RASSF1A (Blood) | RASSF1A (Tissue) | RASSF2A (Blood) | RASSF2A (Tissue) | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Parameters | N | M | U | p | M | U | P | M | U | P | M | U | P Value |
| Over weight + | 19 | 11 | 8 | 0.741 | 11 | 8 | 0.657 | 12 | 7 | 0.945 | 12 | 7 | 0.856 |
| Over weight - | 9 | 5 | 4 | 6 | 3 | 5 | 4 | 6 | 3 | ||||
| Diabetes + | 11 | 9 | 2 | 0.374 | 7 | 4 | 0.576 | 7 | 4 | 0.619 | 7 | 4 | 0.463 |
| Diabetes _ | 17 | 11 | 6 | 9 | 8 | 9 | 8 | 13 | 4 | ||||
| Blood pressure + | 8 | 6 | 2 | 0.186 | 5 | 3 | 0.716 | 5 | 3 | 0.390 | 6 | 2 | 1.0 |
| Blood pressure - | 20 | 12 | 8 | 11 | 9 | 11 | 9 | 15 | 5 |
Relationship Between Clinical Characteristics of the Study Groups with Endometrial Cancer
| Number of Sampeles | Metylation, % | P Value | |
|---|---|---|---|
| Diabetes | 11 | 39.9 | 0.016 |
| Over weight | 19 | 67.9 | 0.026 |
| Blood pressure | 8 | 28.6 | NS |
| Low thyroid | 4 | 18.2 | NS |
| Hypercholesterolemia | 3 | 13.6 | NS |
4. Discussion
There is no effective screening or diagnostic tool for down staging of endometrial cancer, thus its incidence is rising when compared to other gynecological malignancies, (e.g. cervical cancer). In contrast, common risk and predisposing epidemiological and histopathological factors associated with the development of uterine carcinomas have been identified. Based on these factors, selection of the women at higher risk for disease origin and offer them increased attention. Moreover, scientists are still in search for new screening methods toward the aim of detecting premalignant at risk lesions or early stages of the disease (27, 28).
The process of silencing by hypermethylation of promotor region is the most important inactivation mechanism of tumor suppressor genes. Epigenetic inactivation may affect the molecular pathways involved in cell immortalization and transformation, but the silencing can be partially relieved by demethylation of the promoter region. Recently the growing list of genes inactivated by promoter hypermethylation provided an opportunity to examine the epigenetic alteration of multiple cancer related genes in different tumors including endometrial cancer and insights have been developed to understand more deeply about role of these alternations in the diagnosis, treatment and prevention of endometrial cancer (29, 30). Development the field of epigenetic of DNA methylation has several advantages compared to conventional biomarkers such as cytology, RNA or protein derived from tumor evaluation: first, DNA is more stable; secondly when the limited amount of tissue or fluid is available, investigation is possible; thirdly promoter hypermethylation usually happened in discrete CpG islands that minizes regional analysis in comparison with mutations that contains multiple exons, and finally a wide range of body fluids can be studied (31, 32).
Studies have demonstrated that conventional tumor markers in serum, such as carcinoembryonic antigen (CEA), are generally insensitive for screening purposes. Consequently, novel serum biomarkers are clearly needed for the early detection of malignancies. Patients with early and advanced stage cancer have abnormally high levels of circulating DNA in the serum or plasma compared to healthy patients or those with non-malignant diseases (33). Nanogram quantities of DNA circulating in the blood are present in healthy individuals, while cancer patients have an average of 219 ng DNA/mL plasma (10 ~ 1,200 ng/mL plasma) (34). The mechanism surrounding the origin of tumoral DNA that is released into the circulation is poorly understood, but it is assumed that DNA is released during necrosis and/or apoptosis of tumor cells (35). It was reported that genetic and epigenetic alterations in serum DNA (such as point mutation like P53 or Ras, gene amplification, loss of heterozygosity, microsatellite instability, and aberrant methylation) are identical to those found in primary human cancers (36). The presence of gene promoter hypermethylation in the serum and plasma DNA has been demonstrated in patients with cancers of the lung, head and neck, liver, colon, stomach, and breast (34). Also several studies have reported RASSF1A methylation levels in DNA isolated from plasma or serum in the range of 23% to 55% (37). The present data demonstrated that aberrant promoter methylation of RASSF1A and RASSF2A was observed in 65% (P = 0.4948), and 11% (P = 0.09) of blood samples. The serum methylation rate was less than methylation rate in tissues about RASSF2A gene; this is likely due to a loss in extraction and bisulfite conversion, instability, and a high background of normal DNA. Circulatory DNA molecules are easily isolated, simple, non-invasive, and sufficiently sensitive method, so they can serve as a promising biomarker for early diagnostic and prognostic of endometrial cancer screening. Further studies are necessary to confirm the findings in larger samples.
In the present study, experimental evidence indicated that the frequency of hypermethylation of the CpG island in the promoter region of tumor suppressor genes RASSF1A and RASSF2A has close relashipship with carcinogenesis of endometrial cancer. We detected that the patterns of methylation of RASSF1A and RASSF2A genes were 53% and 42% of patient’s tissues, whereas methylation was established in 31%, 9% of normal tissues, respectively. For both genes methylation was significantly associated with endonetrial carcinoma (P = 0.03, P = 0.003).
In a similar study performed by Seeber et al. hypermethylation of RASSF1A gene was analyzed in endometrial carcinomas and it reported 79% methylation positively for the observed gene and significantly higher cumulative methylation index of tumor-suppressor gene in EC type I compared to type II (29). Other studies confirmed this finding: RASSF1A promoter region was reported to be methylated, in average 74% of cases of endometrial cancer patients which was associated with diseass progression (38). In a research conducted by Arafa et al. it was observed that methylated promoter occurred in 74 % of patients and was corrolated with decreased gene expression level (39). It should be noted that aberrant methylation is associated with loss of heterozygosity (LOH) in chromosome 3p that frequently occur in endometrial cancer which is related with development, recurrence and survival of cancer and significantly associated with microsatellite instability in endometrial carcinomas that could block the increasing rate of genetic abnormalities in uterine carcinogenesis (40-42).
Fiolka and et al. have previously reported that frequency of RASSF1A aberrant promoter metyaltion as high as 85.5% and 30% in case and controls respectively. Collectively, privious studies confirmed a high frequency (33% up to 85%) of CpG promoter methylation of the RASSF1A gene in endometrial carcinomas. Moreover, this epigenetic alteration showed different frequencies according to the type of disease, with a higher incidence in endometrioid compared to serous or clear cell carcinomas (38, 40).
Due to variable histopathology, significant association between RASSF1A gene promoter metylation and higher degree of tumor, invasion to biometrial and metastasis to pelivic lymph nodes observed. But not a significant difference between menarche parity, history of oral contraceptive consumption, hormone replacement therapy and smoking among patients and the control group were observed (43). In other reseraches correlation of hypermethylation of CpG islands with clinicopathological parameters (tumor grade, myometrial invasion and nodal involvement) has been demonstrated.
Liao et al. (2008) demonstrated an increased risk for endometrial cancer for patients who had RASSF2A gene promoter methylation 25 of 75 cases (33 %). Also there was an increase in endometrial cancer incidence for elderly patients (40). Hesson et al. (2005) observed similar findings for colon cancer patients (44). Weimeng et al. (2011) analyzed combined methylation pattern of RASSF1A and RASSF2A genes as a diagnostic marker for bladder cancer. They observed methylation in 72% of the cancer cells and only 6% of healthy tissues and confirmed that RASSF1A and RASSF2A genes have a distinct methylation pattern in bladder tumor tissues compared to normal tissues (45).
In conclusion, the present study demonstrated that promoter metyaltion of RASSF1A and RASSF2A genes is a frequent epigenetic event in EC. The results indicated that hypermethylation of these genes was involved in some clinical and pathogenesis of the diseass. Furthurmore, the methylation pattern of these genes in blood samples emphasize that this epigenetic event has the potential to be as a molecular marker for cancer and has digonostic and prognostic values for early carcinogenesis detection in EC. Finally, our data represents a clinical tool for the proper management of the EC.
References
-
1.
Jemal A, Siegel R, Ward E, Hao Y, Xu J, Thun MJ. Cancer statistics, 2009. CA Cancer J Clin. 2009;59(4):225-49. [PubMed ID: 19474385]. https://doi.org/10.3322/caac.20006.
-
2.
Renaud MC, Le T, Le T, Bentley J, Farrell S, Fortier MP, et al. Epidemiology and investigations for suspected endometrial cancer. J Obstet Gynaecol Can. 2013;35(4):380-3. [PubMed ID: 23660050].
-
3.
Lutz JM, Francisci S, Mugno E, Usel M, Pompe-Kirn V, Coebergh JW, et al. Cancer prevalence in Central Europe: the EUROPREVAL Study. Ann Oncol. 2003;14(2):313-22. [PubMed ID: 12562661].
-
4.
Armstrong AJ, Hurd WW, Elguero S, Barker NM, Zanotti KM. Diagnosis and management of endometrial hyperplasia. J Minim Invasive Gynecol. 2012;19(5):562-71. [PubMed ID: 22863972]. https://doi.org/10.1016/j.jmig.2012.05.009.
-
5.
Sorosky JI. Endometrial cancer. Obstet Gynecol. 2012;120(2 Pt 1):383-97. [PubMed ID: 22825101]. https://doi.org/10.1097/AOG.0b013e3182605bf1.
-
6.
Lax S. [Precursor lesions of endometrial carcinoma: diagnostic approach and molecular pathology]. Pathologe. 2011;32 Suppl 2:255-64. [PubMed ID: 22033684]. https://doi.org/10.1007/s00292-011-1514-3.
-
7.
Tao MH, Freudenheim JL. DNA methylation in endometrial cancer. Epigenetics. 2010;5(6):491-8. [PubMed ID: 20543579].
-
8.
Di Domenico M, Santoro A, Ricciardi C, Iaccarino M, Iaccarino S, Freda M, et al. Epigenetic fingerprint in endometrial carcinogenesis: the hypothesis of a uterine field cancerization. Cancer Biol Ther. 2011;12(5):447-57. [PubMed ID: 21709441].
-
9.
Turek-Plewa J, Jagodzinski PP. The role of mammalian DNA methyltransferases in the regulation of gene expression. Cell Mol Biol Lett. 2005;10(4):631-47. [PubMed ID: 16341272].
-
10.
Widschwendter M, Jones PA. DNA methylation and breast carcinogenesis. Oncogene. 2002;21(35):5462-82. [PubMed ID: 12154408]. https://doi.org/10.1038/sj.onc.1205606.
-
11.
Eckfeld K, Hesson L, Vos MD, Bieche I, Latif F, Clark GJ. RASSF4/AD037 is a potential ras effector/tumor suppressor of the RASSF family. Cancer Res. 2004;64(23):8688-93. [PubMed ID: 15574778]. https://doi.org/10.1158/0008-5472.CAN-04-2065.
-
12.
Khokhlatchev A, Rabizadeh S, Xavier R, Nedwidek M, Chen T, Zhang XF, et al. Identification of a novel Ras-regulated proapoptotic pathway. Curr Biol. 2002;12(4):253-65. [PubMed ID: 11864565].
-
13.
Ortiz-Vega S, Khokhlatchev A, Nedwidek M, Zhang XF, Dammann R, Pfeifer GP, et al. The putative tumor suppressor RASSF1A homodimerizes and heterodimerizes with the Ras-GTP binding protein Nore1. Oncogene. 2002;21(9):1381-90. [PubMed ID: 11857081]. https://doi.org/10.1038/sj.onc.1205192.
-
14.
Vos MD, Ellis CA, Elam C, Ulku AS, Taylor BJ, Clark GJ. RASSF2 is a novel K-Ras-specific effector and potential tumor suppressor. J Biol Chem. 2003;278(30):28045-51. [PubMed ID: 12732644]. https://doi.org/10.1074/jbc.M300554200.
-
15.
Song MS, Song SJ, Ayad NG, Chang JS, Lee JH, Hong HK, et al. The tumour suppressor RASSF1A regulates mitosis by inhibiting the APC-Cdc20 complex. Nat Cell Biol. 2004;6(2):129-37. [PubMed ID: 14743218]. https://doi.org/10.1038/ncb1091.
-
16.
Dallol A, Agathanggelou A, Fenton SL, Ahmed-Choudhury J, Hesson L, Vos MD, et al. RASSF1A interacts with microtubule-associated proteins and modulates microtubule dynamics. Cancer Res. 2004;64(12):4112-6. [PubMed ID: 15205320]. https://doi.org/10.1158/0008-5472.CAN-04-0267.
-
17.
Hamilton DW, Lusher ME, Lindsey JC, Ellison DW, Clifford SC. Epigenetic inactivation of the RASSF1A tumour suppressor gene in ependymoma. Cancer Lett. 2005;227(1):75-81. [PubMed ID: 16051033]. https://doi.org/10.1016/j.canlet.2004.11.044.
-
18.
Rykova EY, Laktionov PP, Skvortsova TE, Starikov AV, Kuznetsova NP, Vlassov VV. Extracellular DNA in breast cancer: Cell-surface-bound, tumor-derived extracellular DNA in blood of patients with breast cancer and nonmalignant tumors. Ann N Y Acad Sci. 2004;1022:217-20. [PubMed ID: 15251963]. https://doi.org/10.1196/annals.1318.033.
-
19.
Thaler S, Hahnel PS, Schad A, Dammann R, Schuler M. RASSF1A mediates p21Cip1/Waf1-dependent cell cycle arrest and senescence through modulation of the Raf-MEK-ERK pathway and inhibition of Akt. Cancer Res. 2009;69(5):1748-57. [PubMed ID: 19223555]. https://doi.org/10.1158/0008-5472.CAN-08-1377.
-
20.
Song MS, Song SJ, Kim SY, Oh HJ, Lim DS. The tumour suppressor RASSF1A promotes MDM2 self-ubiquitination by disrupting the MDM2-DAXX-HAUSP complex. EMBO J. 2008;27(13):1863-74. [PubMed ID: 18566590]. https://doi.org/10.1038/emboj.2008.115.
-
21.
Hesson L, Bieche I, Krex D, Criniere E, Hoang-Xuan K, Maher ER, et al. Frequent epigenetic inactivation of RASSF1A and BLU genes located within the critical 3p21.3 region in gliomas. Oncogene. 2004;23(13):2408-19. [PubMed ID: 14743209]. https://doi.org/10.1038/sj.onc.1207407.
-
22.
Hesson LB, Wilson R, Morton D, Adams C, Walker M, Maher ER, et al. CpG island promoter hypermethylation of a novel Ras-effector gene RASSF2A is an early event in colon carcinogenesis and correlates inversely with K-ras mutations. Oncogene. 2005;24(24):3987-94. [PubMed ID: 15806169]. https://doi.org/10.1038/sj.onc.1208566.
-
23.
Cooper WN, Hesson LB, Matallanas D, Dallol A, von Kriegsheim A, Ward R, et al. RASSF2 associates with and stabilizes the proapoptotic kinase MST2. Oncogene. 2009;28(33):2988-98. [PubMed ID: 19525978]. https://doi.org/10.1038/onc.2009.152.
-
24.
Richter AM, Pfeifer GP, Dammann RH. The RASSF proteins in cancer; from epigenetic silencing to functional characterization. Biochim Biophys Acta. 2009;1796(2):114-28. [PubMed ID: 19344752]. https://doi.org/10.1016/j.bbcan.2009.03.004.
-
25.
Wang Y, Yu Z, Wang T, Zhang J, Hong L, Chen L. Identification of epigenetic aberrant promoter methylation of RASSF1A in serum DNA and its clinicopathological significance in lung cancer. Lung Cancer. 2007;56(2):289-94. [PubMed ID: 17267069]. https://doi.org/10.1016/j.lungcan.2006.12.007.
-
26.
Song H, Oh S, Oh HJ, Lim DS. Role of the tumor suppressor RASSF2 in regulation of MST1 kinase activity. Biochem Biophys Res Commun. 2010;391(1):969-73. [PubMed ID: 19962960]. https://doi.org/10.1016/j.bbrc.2009.11.175.
-
27.
Heichman KA, Warren JD. DNA methylation biomarkers and their utility for solid cancer diagnostics. Clin Chem Lab Med. 2012;50(10):1707-21. [PubMed ID: 23089699]. https://doi.org/10.1515/cclm-2011-0935.
-
28.
Hatzimichael E, Crook T. Cancer epigenetics: new therapies and new challenges. J Drug Deliv. 2013;2013:529312. [PubMed ID: 23533770]. https://doi.org/10.1155/2013/529312.
-
29.
Pallares J, Velasco A, Eritja N, Santacana M, Dolcet X, Cuatrecasas M, et al. Promoter hypermethylation and reduced expression of RASSF1A are frequent molecular alterations of endometrial carcinoma. Mod Pathol. 2008;21(6):691-9. [PubMed ID: 18469797]. https://doi.org/10.1038/modpathol.2008.38.
-
30.
Foley DL, Craig JM, Morley R, Olsson CA, Dwyer T, Smith K, et al. Prospects for epigenetic epidemiology. Am J Epidemiol. 2009;169(4):389-400. [PubMed ID: 19139055]. https://doi.org/10.1093/aje/kwn380.
-
31.
Qian ZR, Sano T, Yoshimoto K, Yamada S, Ishizuka A, Mizusawa N, et al. Inactivation of RASSF1A tumor suppressor gene by aberrant promoter hypermethylation in human pituitary adenomas. Lab Invest. 2005;85(4):464-73. [PubMed ID: 15711568]. https://doi.org/10.1038/labinvest.3700248.
-
32.
Pan ZG, Kashuba VI, Liu XQ, Shao JY, Zhang RH, Jiang JH, et al. High frequency somatic mutations in RASSF1A in nasopharyngeal carcinoma. Cancer Biol Ther. 2005;4(10):1116-22. [PubMed ID: 16096369].
-
33.
Matthaios D, Balgkouranidou I, Karayiannakis A, Bolanaki H, Xenidis N, Amarantidis K, et al. Methylation status of the APC and RASSF1A promoter in cell-free circulating DNA and its prognostic role in patients with colorectal cancer. Oncol Lett. 2016;12(1):748-56. [PubMed ID: 27347211]. https://doi.org/10.3892/ol.2016.4649.
-
34.
Bae YK, Shim YR, Choi JH, Kim MJ, Gabrielson E, Lee SJ, et al. Gene promoter hypermethylation in tumors and plasma of breast cancer patients. Cancer Res Treat. 2005;37(4):233-40. [PubMed ID: 19956520]. https://doi.org/10.4143/crt.2005.37.4.233.
-
35.
Jahr S, Hentze H, Englisch S, Hardt D, Fackelmayer FO, Hesch RD, et al. DNA fragments in the blood plasma of cancer patients: quantitations and evidence for their origin from apoptotic and necrotic cells. Cancer Res. 2001;61(4):1659-65. [PubMed ID: 11245480].
-
36.
Wang YC, Yu ZH, Liu C, Xu LZ, Yu W, Lu J, et al. Detection of RASSF1A promoter hypermethylation in serum from gastric and colorectal adenocarcinoma patients. World J Gastroenterol. 2008;14(19):3074-80. [PubMed ID: 18494062].
-
37.
Yazici H, Terry MB, Cho YH, Senie RT, Liao Y, Andrulis I, et al. Aberrant methylation of RASSF1A in plasma DNA before breast cancer diagnosis in the Breast Cancer Family Registry. Cancer Epidemiol Biomarkers Prev. 2009;18(10):2723-5. [PubMed ID: 19755643]. https://doi.org/10.1158/1055-9965.EPI-08-1237.
-
38.
Seeber LM, Zweemer RP, Marchionni L, Massuger LF, Smit VT, van Baal WM, et al. Methylation profiles of endometrioid and serous endometrial cancers. Endocr Relat Cancer. 2010;17(3):663-73. [PubMed ID: 20488783]. https://doi.org/10.1677/ERC-10-0014.
-
39.
Arafa M, Kridelka F, Mathias V, Vanbellinghen JF, Renard I, Foidart JM, et al. High frequency of RASSF1A and RARb2 gene promoter methylation in morphologically normal endometrium adjacent to endometrioid adenocarcinoma. Histopathology. 2008;53(5):525-32. [PubMed ID: 18783461]. https://doi.org/10.1111/j.1365-2559.2008.03147.x.
-
40.
Liao X, Siu MK, Chan KY, Wong ES, Ngan HY, Chan QK, et al. Hypermethylation of RAS effector related genes and DNA methyltransferase 1 expression in endometrial carcinogenesis. Int J Cancer. 2008;123(2):296-302. [PubMed ID: 18404674]. https://doi.org/10.1002/ijc.23494.
-
41.
Kang S, Lee JM, Jeon ES, Lee S, Kim H, Kim HS, et al. RASSF1A hypermethylation and its inverse correlation with BRAF and/or KRAS mutations in MSI-associated endometrial carcinoma. Int J Cancer. 2006;119(6):1316-21. [PubMed ID: 16619251]. https://doi.org/10.1002/ijc.21991.
-
42.
Kang S, Kim JW, Kang GH, Lee S, Park NH, Song YS. Comparison of DNA hypermethylationpatterns in different types of uterine cancer, cervicalsquamous cell carcinoma, cervical adenocarcinoma andendometrial adenocarcinoma. Int J Cancer. 2006;118:2168-71.
-
43.
Fiolka R, Zubor P, Janusicova V, Visnovsky J, Mendelova A, Kajo K, et al. Promoter hypermethylation of the tumor-suppressor genes RASSF1A, GSTP1 and CDH1 in endometrial cancer. Oncol Rep. 2013;30(6):2878-86. [PubMed ID: 24068440]. https://doi.org/10.3892/or.2013.2752.
-
44.
Hesson LB, Latif F. RASSF2 (Ras association (RalGDS/AF-6) domain family member 2). Oncol Haematol. 2010;14(7):652-61.
-
45.
Meng W, Huebner A, Shabsigh A, Chakravarti A, Lautenschlaeger T. Combined RASSF1A and RASSF2A Promoter Methylation Analysis as Diagnostic Biomarker for Bladder Cancer. Mol Biol Int. 2012;2012:701814. [PubMed ID: 22530128]. https://doi.org/10.1155/2012/701814.